castanyes blaves
Random ramblings about some random stuff, and things; but more stuff than things -- all in a mesmerizing and kaleidoscopic soapbox-like flow of words.9/30/2008
A supercomputing architecture that first appeared in prototype form more than 10 years ago has been given a new lease on life, thanks in part to a recent $4 million Department of Defense grant issued to seed the new Center for Adaptive Supercomputing Software. The joint project teams up Pacific Northwest National Laboratory and supercomputer maker Cray, as well as several institutions including Georgia Institute of Technology and Sandia National Laboratories.
The initiative aims to develop software that takes advantage of the multithreaded processing capabilities of Cray's XMT supercomputer.
Dubbed Magalhaes (Magellan), the laptops will have on board low-power Intel Atom chips designed for laptops. They will also sport digital cameras and a broadband net connection.
As an operating system, the machines will run a version of Linux developed in Venezuela.
9/23/2008
On 23 September the US arm of operator T-Mobile is expected to whisk the cloth off the first handset running Google's Android operating system for mobiles.
VMware Fusion 2.0 goes final: free update to existing users
Report: Apple now sixth among worldwide PC manufacturers
By David Chartier | Published: September 13, 2008 - 12:08PM CT
Riding the wave of good news about Apple's explosive growth over the past years, market research firm Gartner now says that Apple is the sixth largest PC manufacturer in the world.
Challenge to iPhone heats up
By Andrew Parker in London and Paul Taylor in New York
Published: September 22 2008 23:30 | Last updated: September 22 2008 23:30
The battle to dominate the mobile smartphone market is about to heat up with the launch of long-awaited handsets by Google, Nokia and Sony Ericsson.
On Tuesday, Google is due to unveil the first smartphone powered by the internet search company’s Android operating system.
9/18/2008
Code size open source projects
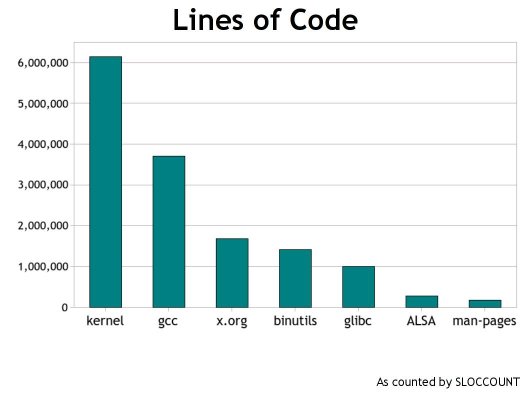
9/17/2008
Fast spores
Escaping the dung pile quickly: Speedy Pilobolus spores
This video is really cool:
Video S4
9/10/2008
Scientists have hailed a successful switch-on for an enormous experiment which will recreate the conditions a few moments after the Big Bang.
9/08/2008
Indels and mutation
Single-nucleotide mutation rate increases close to insertions/deletions in eukaryotes
Very simple but elegant analysis. They used different alignment programs to check for errors, and the plots show distances from 0 to 25bp and 0 to 50bp. Say an indel is at 12bp of a mismatch: are we properly aligning the sequences? Would PRANK give different results on a set of human-chimp-orang-macaque multiple alignments compared to the pairwise alignments methods used in the paper? The results indicate that this is not a recombination effect. That's interesting. There is a pile of papers out there that relate polymorphism (and divergence?) to recombination rates. Not exactly contradictory, but interesting to see the difference in results.
Phylogenetic Inference Using Whole Genomes
Phylogenetic Inference Using Whole Genomes
Bruce Rannala1 and Ziheng Yang2
1Genome Center and Department of Evolution and Ecology, University of California, Davis, California 95616; email: bhrannala@ucdavis.edu
2Department of Biology, University College London, London WC1E 6BT United Kingdom; Laboratory of Biometrics, Graduate School of Agriculture and Life Sciences, University of Tokyo, Tokyo, Japan; email: z.yang@ucl.ac.uk
The availability of genome-wide data provides unprecedented opportunities for resolving difficult phylogenetic relationships and for studying population genetic processes of mutation, selection, and recombination on a genomic scale. The use of appropriate statistical models becomes increasingly important when we are faced with very large datasets, which can lead to improved precision but not necessarily improved accuracy if the analytical methods have systematic biases. This review provides a critical examination of methods for analyzing genomic datasets from multiple loci, including concatenation, separate gene-by-gene analyses, and statistical models that accommodate heterogeneity in different aspects of the evolutionary process among data partitions. We discuss factors that may cause the gene tree to differ from the species tree, as well as strategies for estimating species phylogenies in the presence of gene tree conflicts. Genomic datasets provide computational and statistical challenges that are likely to be a focus of research for years to come.
Interesting read. The supertree (separate model) method seems to be a dangerous option in certain cases. The performance can even deteriorate with the inclusion of more genes in the dataset.
Archives
200409 200412 200501 200502 200503 200504 200505 200506 200507 200508 200509 200510 200511 200512 200601 200602 200603 200604 200605 200606 200607 200608 200609 200610 200611 200612 200701 200702 200703 200704 200705 200707 200708 200709 200710 200711 200712 200801 200802 200803 200804 200805 200806 200807 200808 200809 200810 200811 200812 200901 200902 200903 200904 200905 200906 200907 200908 200909 200912 201001 201002 201003 201004 201007 201009 201011 201102
Subscribe to Comments [Atom]